- Details
Inbal Mermershtian from the Kosloff lab used a computational structural bioinformatics pipeline that requires hundreds of cores working in parallel for accurate atomic-level calculations of protein-protein interactions - analyzing multiple proteins 3D complexes quantitatively.
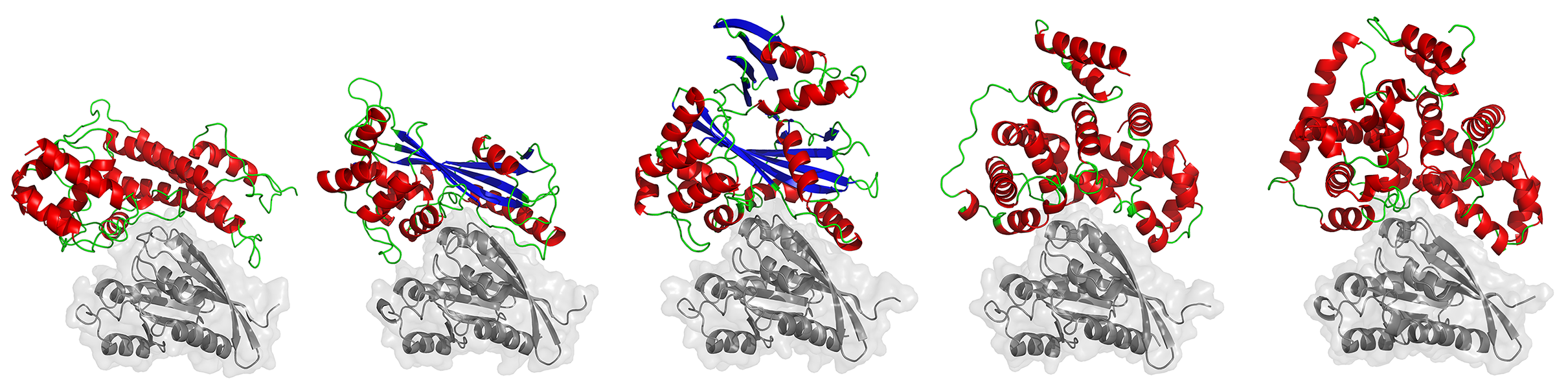
In this project Inbal focused on the small G-proteins Rabs, comparing their modulation by host GAP proteins (that turn Rab switches “off”) with the modulation by pathogen GAP proteins. She use a computational method developed in the Kosloff lab that applies structure-based electrostatic energy calculations to pinpoint structural determinants that are critical for fine-tuning protein-protein interaction specificity.
Inbal's analysis uncovers a convergent structural basis for Rab recognition and modulation by eukaryotic and bacterial pathogens, and highlighted key specificity-determining interactions that might be used to target such pathogens with new antibiotics. More generally, this study presents a fascinating case-study of how similar interaction specificity can be achieved by structurally-dissimilar proteins.
- Details
 Rana Saad from the Privman lab came first place in Hive usage in 2015! She used 698,830 CPU hours, out of a total of 1,135,463 CPU hours used on the cluster in 2015.
Rana Saad from the Privman lab came first place in Hive usage in 2015! She used 698,830 CPU hours, out of a total of 1,135,463 CPU hours used on the cluster in 2015.
Page 2 of 2
